Web Browser Requirements
inpPDH requires a modern web browser with JavaScript and cookies enabled. To view the complex details, pop-ups must not be blocked. The following browsers have been thoroughly tested with inpPDH:
- Mozilla Firefox, version 4 or above
- Internet Explorer, versions 11 or above
- Chrome, version 5 or above
Usage
1. Input
Two types of input formats are allowed for inpPDH: protein-DNA structural files from PDB (Protein Data Bank) or PDB IDs. As there are more than one protein chain in protein-DNA complexes, chain information is also needed as input data. Submission title and email address are optional for users to fill.Notes:
- Only a protein-DNA complex with interactional interface is allowed for prediction.
- inpPDH is not able to identify the structure with several models. We only keep the first model of structure for calculation. If you want to calculate a certain model specifically, please make a process of submitted PDB file first.
- The modified non-standard residues labeled as HETATM cannot be read by inpPDH.
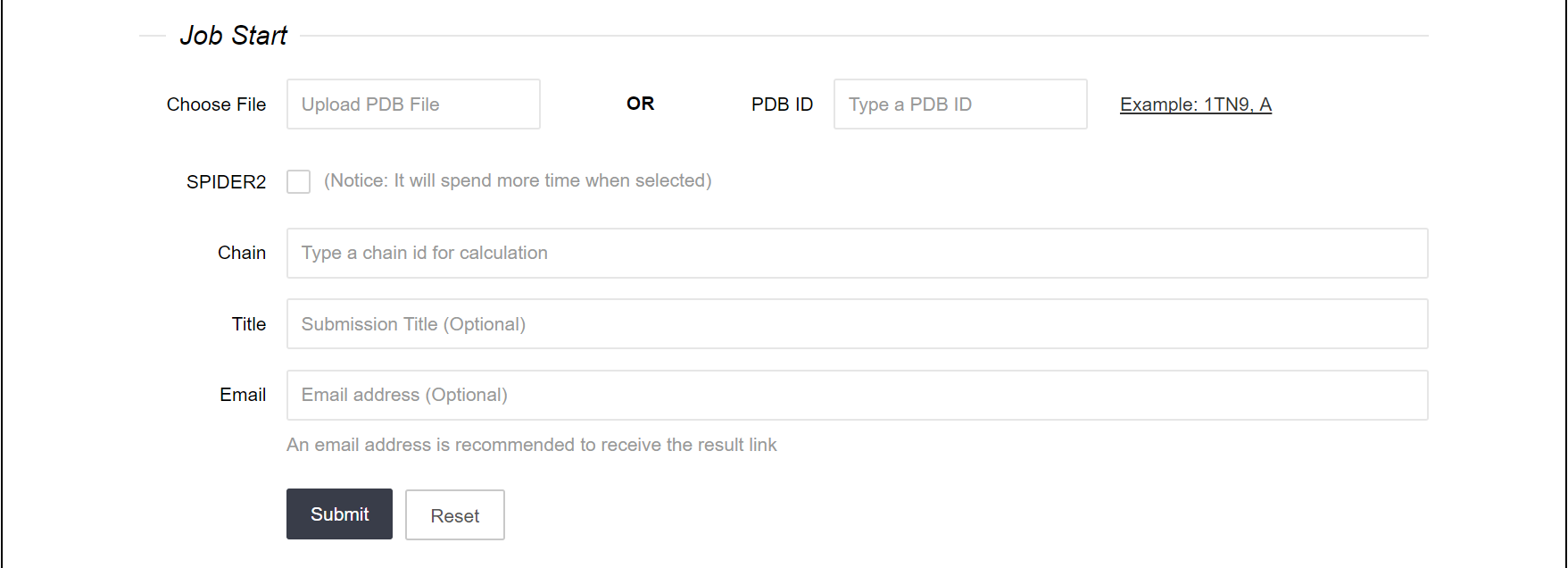
Figure 1.
2. Submission
inpPDH runs tasks one by one. Multiple submitted tasks are queued for execution. If you submit the job mistakenly, the "Cancel" button is prepared for you to terminate the job in queue. Only the job in the queue can be canceled. Once the job is submitted, a table containing job information (job ID, created time, execution status and so on) can be returned. The result link is shown on the web page or sent to you (if you leave the email address).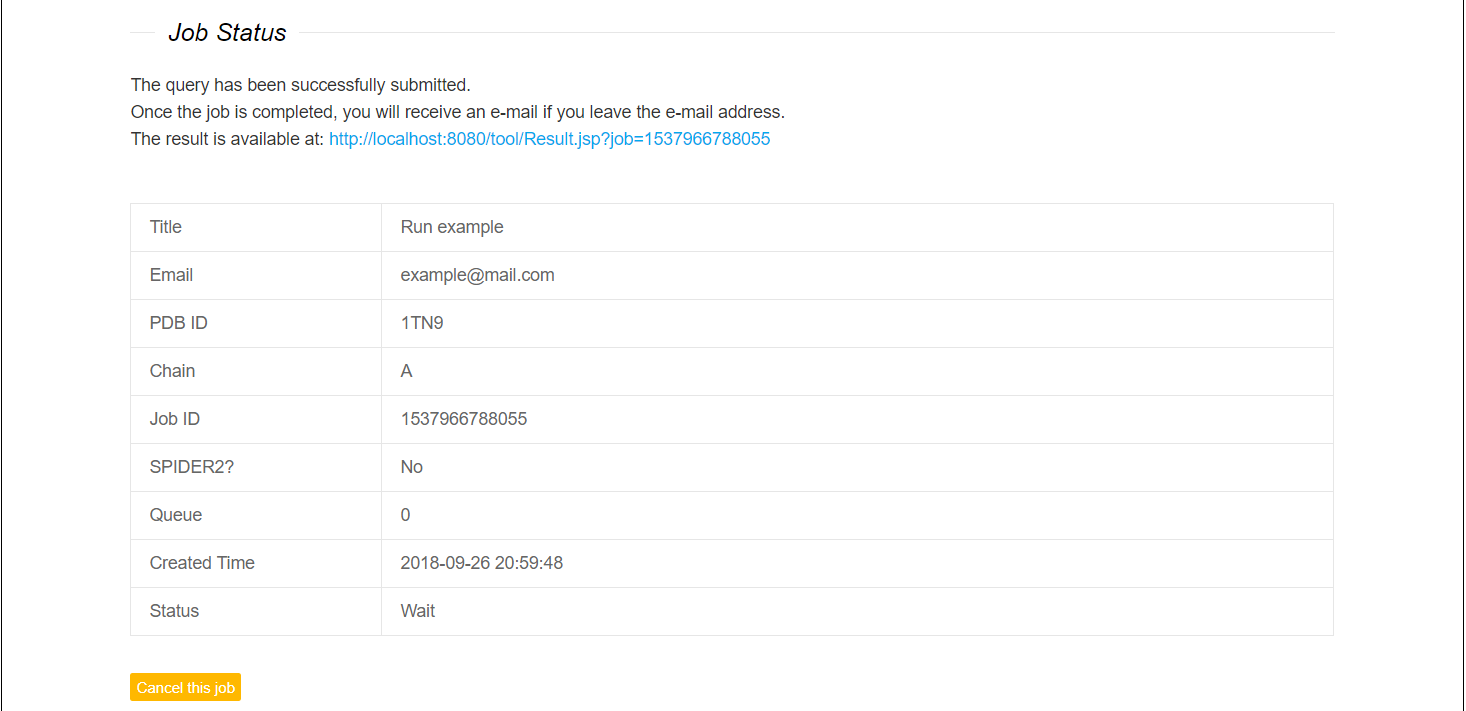
3. Output
The predicted results will be displayed in a table. Meanwhile, we provide users with a visualized three-dimensional structure of protein-DNA complexes. It is executable to mark corresponding positions of predicted hot spots and non-hot spots on the visual structures. All prediction results can be download in tar.gz form.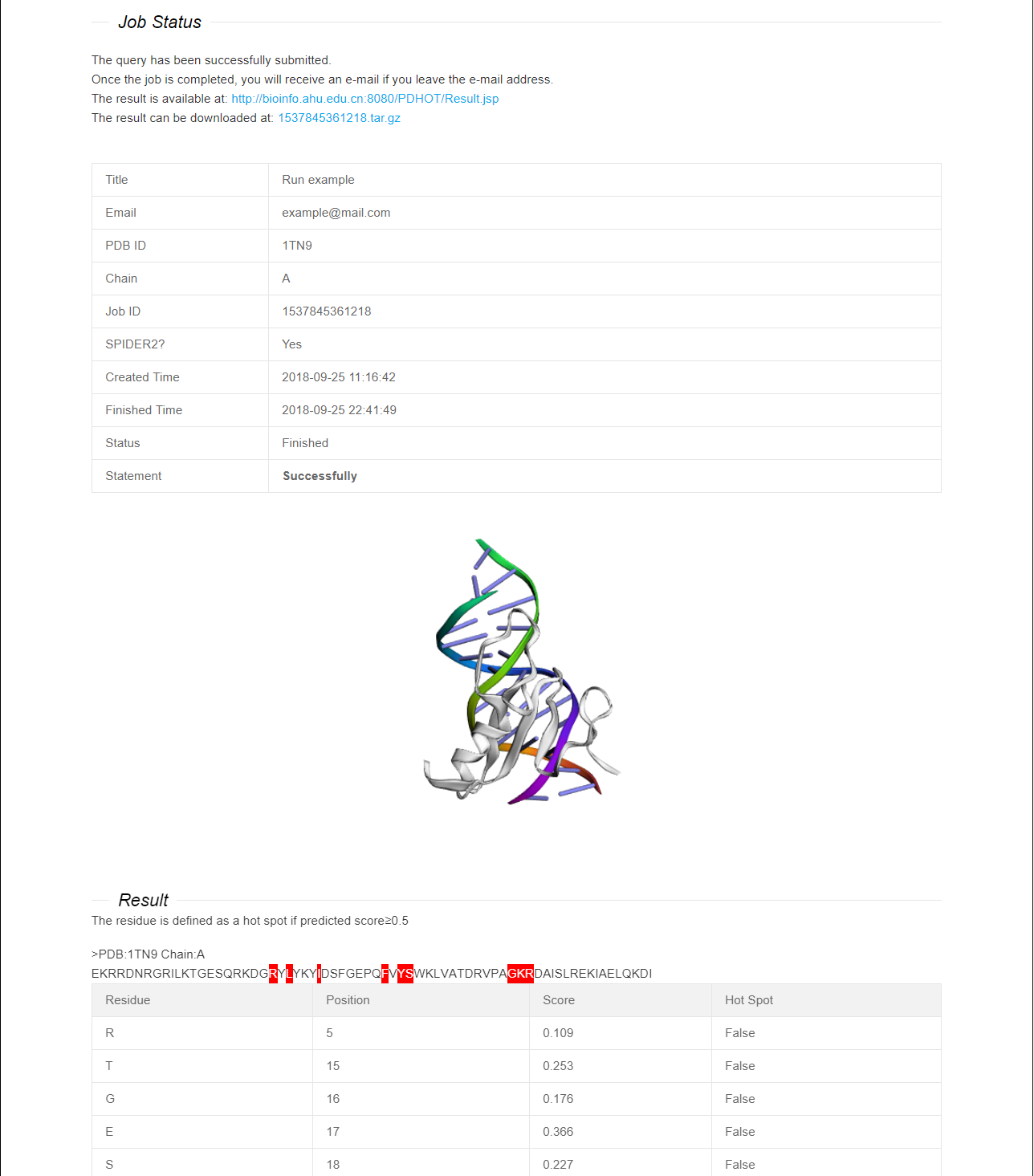
4. Example
The example on the Server page provides a PDB ID (1TN9) and its chain (A) information. If you want to upload an example of PDB file, we also offer a download link here 1TN9.pdb. Upload the file and fill in the blank of chain "A". You can also use job ID and left email address to search for the task on the Server page. An example job ID is given.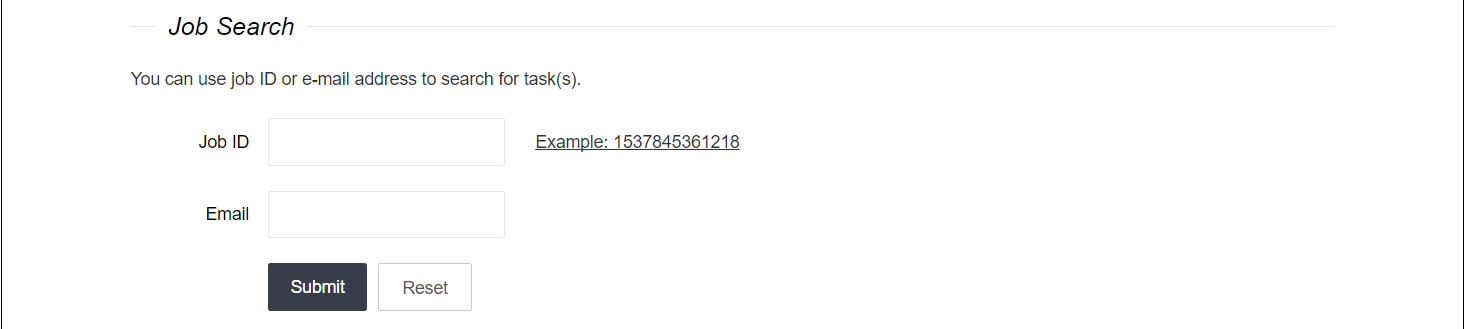
Version
The current version number is 2.0 - Nov, 2019.